|
8/19/2023 0 Comments 4peaks software download windows 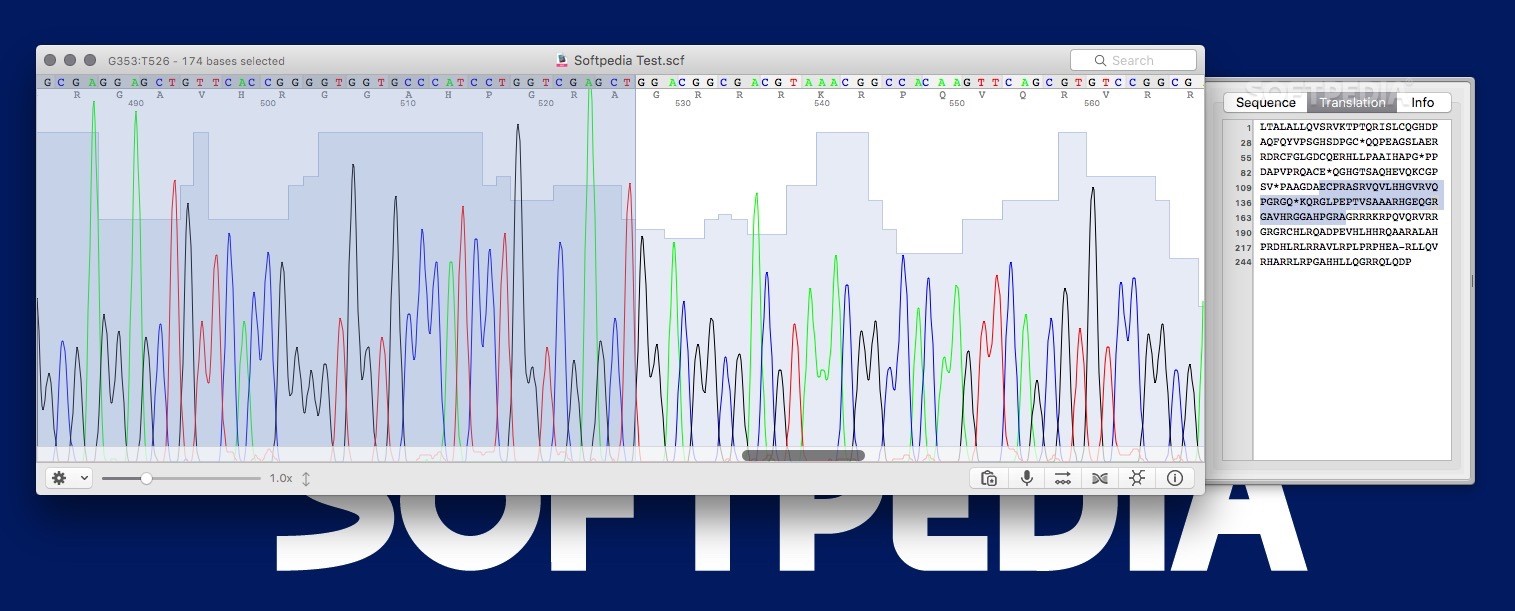
Select the good-quality region using multiple alignments with high-quality reference sequences and manual inspection of the chromatogram.Use only a good-quality region of your sequence.The example below looks OK at a first glance, but this region should be trimmed. In the worst cases, you will need to sequence again. A Case of bad qualityĪny region of low-quality chromatogram should be trimmed. Even though there are cases where the different species show identical 16S sequences, this case clearly indicates that the isolate belongs to Ruminococcus faecis. (a) After trimming, your sequence is 100% identical to the hit ( Ruminococcus faecis). Paste to the EzBioCloud “Identify” as a new query. To copy your trimmed sequence, (a) select “Select Panel” and (b) select “Copy as FASTA format”. The trimmed sequence can be obtained here as FASTA, EzEditor2 file. I trimmed at the absolute position of 1,213 which left 519 bases. In this example, the ab1 contains 1,539 bases (very long for a single Sanger reaction). 400 bp is sufficient to get an accurate identification. Find the position where you want to trim the right-side (the rest of sequence). Compare your sequence with the hit reference sequence and also consult the chromatogram. This will move the cursor to the next position where nucleotide does not match to the hit reference sequence. A tip is to use the shortcut Ctrl+U key (Compare UP function). Right-click to activate the popup menu then select “Delete”->”Before cursor”. Move the cursor to the position where you want to trim. You need to trim this erroneous region in the EzEditor2 tool. The corresponding region in the chromatogram also shows low quality sequencing reaction (box (b) of the blow). The sequence of the best hit ( Ruminococcus faecis) is located at the right upper row of your sequence).Īlignment your sequence against the hit sequence ( Ruminococcus faecis) by clicking (c).Īfter the automated pairwise alignment, you will see the erroneous region that does not match to other reference sequences (box (a) of the above). Check your sequence (from ab1 file) is at the bottom (b). Move the cursor (a) to the bottom of the screen. Your sequence is located at the bottom of the alignment. Start EzEditor2 software and open the file that you have just downloaded. This file contains your sequences and reference sequences that show the highest sequence similarities. Click (c) to view the detailed result of “Identify”.Ĭlick (a) to download a data file for EzEditor2 tool. We will start the trimming process using a sequence alignment tool called EzEditor2. As you will see the chromatogram shown above, your sequence contains errors at the front end (as well as backend). This is cutoff is applicable for high-quality sequence only. The generally accepted 16S similarity cutoff is 98.7%. In this case, sequence similarity is 95.35% and you may be excited to discover a potentially new species. The hit species (a) is displayed with pairwise sequence similarity (b). Go to and paste your sequence as new query. Then, search through the EzBioCloud’s Identify. This file contains the result of the fairly good sequencing reaction. This contains a 16S sequence of a human fecal bacterium. If you used the sequence data without trimming the ends, your sequence will have a substantial amount of errors. Because of the sequencing chemistry, both ends of sequence contain substantial errors. Quality and manual inspection of the raw data is possible with the chromatogram. An ab1 file contains both the chromatogram and sequence data (after base calling).The following example ab1 files are required.Ĭharacteristics of Sanger sequencing data You need a sequence alignment tool for 16S gene sequences.If your sequencing service provider did not give you ab1 files, ask for ab1 files. An ab1 file contains the chromatogram information from ABI sequencers ( Learn more). This document explains how to use EzBioCloud’s “Identity” service using single Sanger sequencing data.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
